|
5/8/2023 0 Comments Messenger rnaWe describe what is known about how miRNAs target mRNAs for rapid decay and translation repression, and highlight recent studies that have begun to pinpoint how miRNAs inhibit translation initiation. We discuss two distinct models for how NMD distinguishes between normal and aberrant PTC-bearing mRNAs, and suggest ways that they can be reconciled via a ‘unified' model. Remodeling events are likely to be crucial for both miRNA-mediated silencing and NMD ( Schell et al, 2002 Dreyfuss et al, 2003 Maquat, 2004 Amrani et al, 2006 Chang et al, 2007 Jackson and Standart, 2007 Nilsen, 2007 Pillai et al, 2007). The other is microRNA (miRNA)-mediated silencing of gene expression, which involves the base pairing of miRNAs with the 3′ untranslated regions (UTRs) of their target mRNAs. One is nonsense-mediated mRNA decay (NMD), an RNA surveillance mechanism that rapidly degrades mRNAs harboring premature termination codons (PTCs). In this review, we use two regulatory mechanisms that control mRNA translation and decay as examples to illustrate how a decision may be reached to translate or to degrade a cytoplasmic mRNA. Thus, mRNP remodeling is likely to play a critical role in forming decision as to whether to translate or to degrade an mRNA. Individual mRNA–protein complex (mRNP) components may serve as adaptors that allow mRNAs to interface with the machinery mediating their subcellular localization, translation, and decay. How are these decisions made? Throughout their lifetime, mRNAs associate with a host of proteins factors, some of which are stably bound while others subject to dynamic exchange ( Moore, 2005). 
All mRNAs are ultimately degraded at a defined rate. mRNAs that are initially translated may later be temporarily translationally repressed. Once mRNAs enter the cytoplasm, they are translated, stored for later translation, or degraded. Messenger RNA (mRNA) mediates the transfer of genetic information from the cell nucleus to ribosomes in the cytoplasm, where it serves as a template for protein synthesis. Possible modes involving 3′ untranslated region and its associated factors, which appear to play key roles in both processes, are discussed. In addition, both are linked to RNA processing bodies. 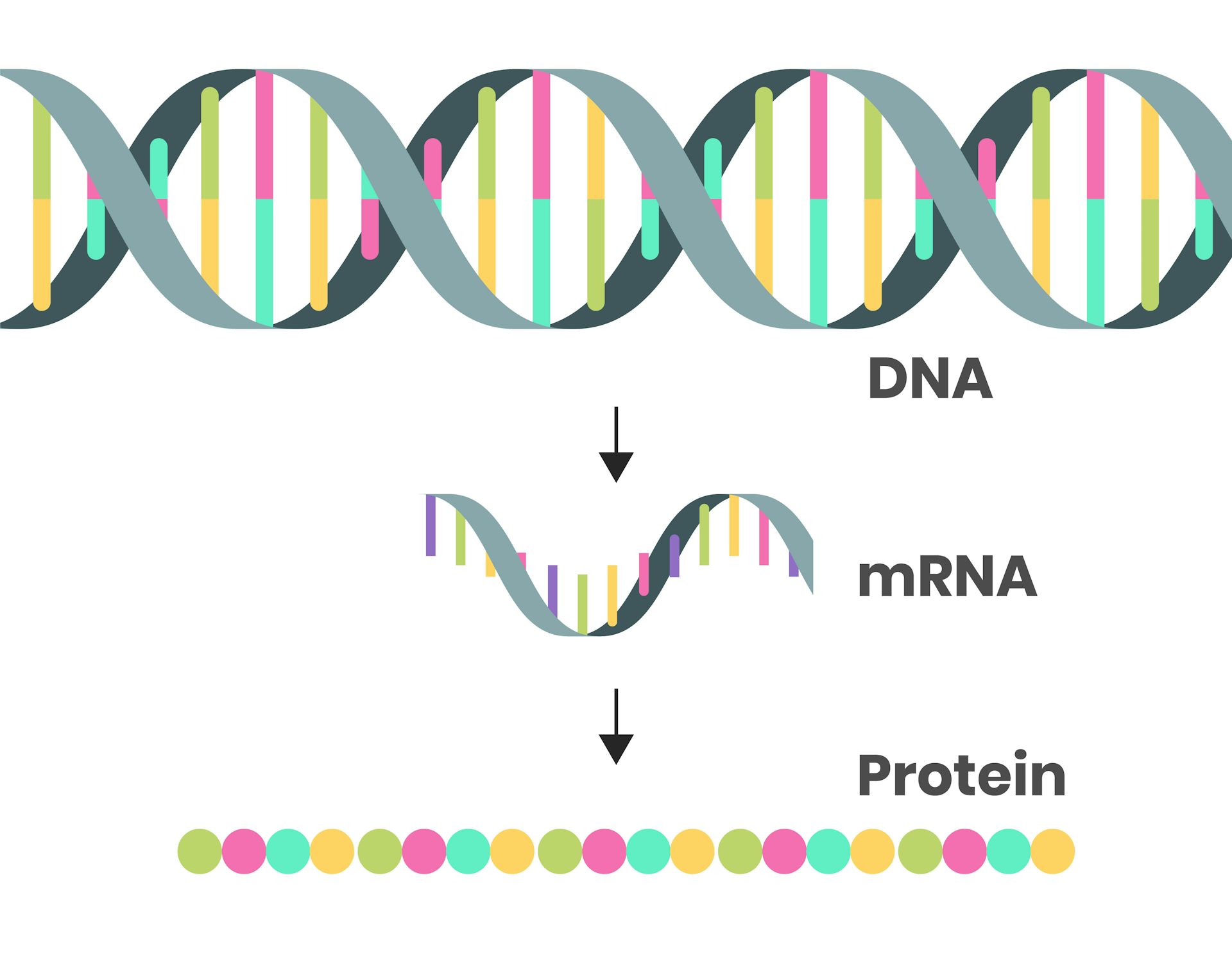
While NMD and miRNA-mediated mRNA silencing use different decision-making processes to determine the fate of their targets, both are greatly influenced by mRNP dynamics. miRNA-mediated mRNA silencing is a mechanism that ensures a given protein is expressed at a proper level to permit normal cellular function. NMD is a surveillance mechanism that detects and eliminates aberrant mRNAs whose expression would result in truncated proteins that are often deleterious to the organism. Nonsense-mediated mRNA decay (NMD) and microRNA (miRNA)-mediated mRNA silencing are provided as examples. In this review we discuss how a decision to translate or to degrade a cytoplasmic mRNA is reached. These mRNA–protein complexes (mRNPs) undergo a series of remodeling events that are influenced by and/or influence the translation and mRNA decay machinery. 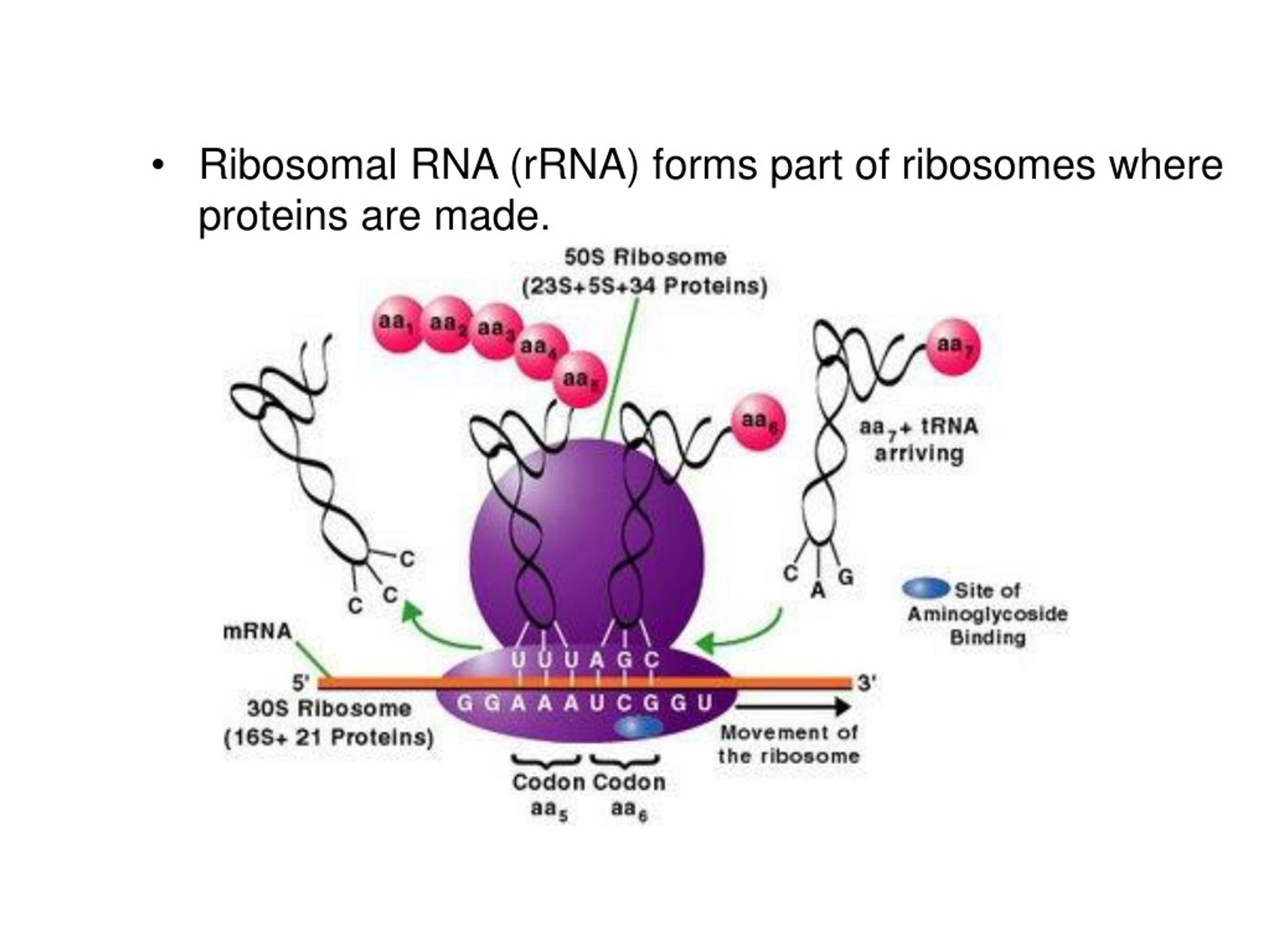 
Once transcribed, mRNAs associate with a host of proteins throughout their lifetime. Quality control of gene expression operates post-transcriptionally at various levels in eukaryotes.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
